Combat-TB-NeoDB

Fostering Tuberculosis research through integrative analysis using graph database technologies.
Combat-TB-NeoDB is an integrated M.tuberculosis (M.tb) ‘omics’ knowledge-base. NeoDB is based on Neo4j and enables researchers to execute complex federated queries by linking well-known, curated and widely used M.tb data resources, and supplementary Tuberculosis variants data from published literature. Combat-TB-NeoDB was created by binding the labeled property graph model to a consensus-controlled ontology.
A publicly accessible instance is available at: https://neodb.sanbi.ac.za
Purpose
To enable M.tuberculosis (M.tb) researchers to execute ad hoc complex federated queries with the ability to explore data interactively by linking well-known, curated and widely used biological data resources, and supplementary TB variants data from published literature.
Installation
To avoid the trouble of environment setup, running the database in a Docker container is highly recommended.
Otherwise, a manual installation guide can be found here.
Graph Model (Schema)
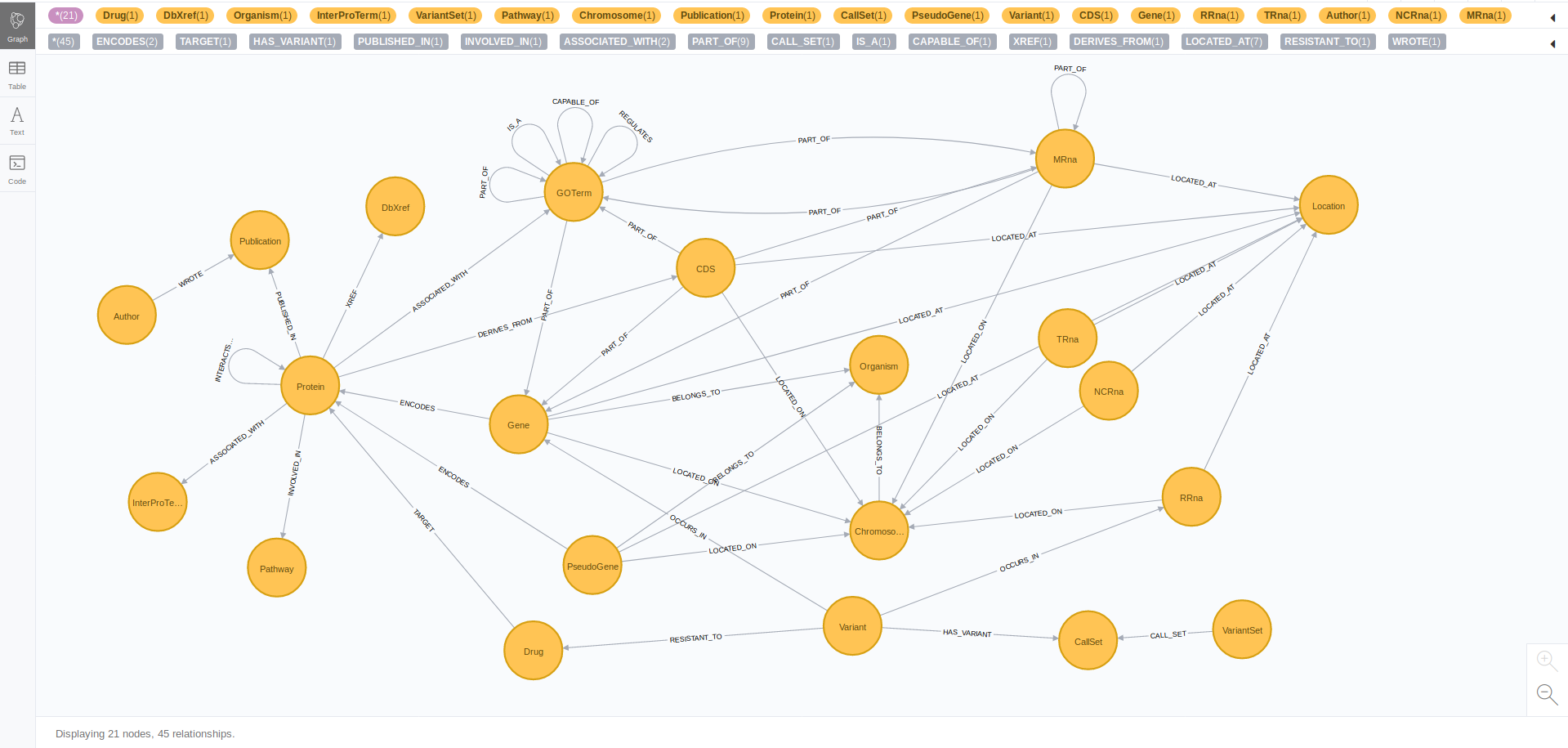
Issues
Please report any issues here.